Optogenetics, the Archons, politics and 'writing circuitry for cells' (transhumanism is now official: 12th Sep 2022), new human blood issues. PART 1.
Note for all email readers: due to new substack size restrictions, the complete version of the post can be found at, that’s here:)
Please stay tuned to continuation in PART 2. Some updates first, the main post starts below the divider bar.
UPDATES FIST, the main post starts below the divider bar below.
10/11/2022: GRAPHENE DECOMPOSITION note, at the very end of this post.
10/6/2022: COVID crimes explained in an article titled “U.S. D.O.D issued a contract for COVID-19 Research to a company in Ukraine, 3 months before COVID-19 was known to exist“ at: https://expose-news.com/2022/10/06/2019-contract-covid-resarch-ukraine-us-dod/ meaning Dr.’s Ralf Baric, Dr. Fauci were not alone in their goals.
10/5/2022: There is nothing better to observe than who is getting Nobel Prize for what. Today the prestigious committee announced a new set of criminals “Nobel Prize for 3 chemists who made molecules ‘click’ “for work on click chemistry and bioorthogonal reactions” with the one quote from: https://apnews.com/article/science-health-stockholm-chemistry-fd3521c6436c94fd6dd73f4e53d86d09 “for developing a way of “snapping molecules together” that can be used to explore cells, map DNA and design drugs that can target diseases such as cancer more precisely.“
10/5/2022: “Scientists Have Discovered a New Set of Blood Groups. The ‘Er’ grouping could help doctors identify and treat some rare cases of blood incompatibility, including between pregnant mothers and fetuses.“!!!???? How interesting, that all the essential changes in biology, are starting to appear first in ‘covid era’, AFTER the covid GENETICALLY modifying injections!!! The link is at: https://www.wired.com/story/new-blood-types/ Essential with that summary is the cited publication tilted: ”Missense mutations in PIEZO1, encoding the Piezo1 mechanosensor protein, define the Er red blood cell antigens” There is no protein with that name yet in uniprot, but the ‘old world version’ human PIEZ1 has very similar W(trp)-motifs which can be found in Dr. Walensky’s 2008 patent for HIV, ebola, influenza, SARS cure… These are the motifs like: ‘‘PPRRQWWRPW’, ‘DWVWTDTTLSLSSWM', or ‘LWRKLLKAFWWL’. Interesting Engineering at https://interestingengineering.com/science/scientists-discover-new-set-of-blood-groups writes about that new discovery, quote: “Genetic variations that code for Piezo1, a protein linked to the new blood system, perceive blood cells as 'foreign’”
Btw. are the in the previous post mentioned transplantants rejections possibly BECAUSE of the changing HUMAN BLOOD in the covid injected victims??? Few more essential remarks about PIEZO1 are at the end of this post.
10/4/2022: The NIH data bank seems to not care about details any more. When you search for human proteins homolog to SARS-CoV-2 Spike (YP_009724390.1) you get the first hit for a human protein(!): Spike glycoprotein [Homo sapiens] linking it to the Sequence ID: 7TLZ_J. That PdbDataBank entry defines the 3D coordinate of that protein coming from: Organism(s): Homo sapiens with the Expression System: Homo sapiens!!??
Title of that work:”Structural basis of SARS-CoV-2 Omicron immune evasion and receptor engagement”. Looks like Omicron lost the furin site completely(!?), no more N-terminal signal, and the entire HIV-1 infectious homolog section at the C-term is gone too… Oh, maybe because now “CDC confirms USA suffered 338x increase in reports of AIDS-associated Diseases & Cancers in 2021 following COVID Vaccine roll-out“ at : https://expose-news.com/2022/10/05/cdc-338x-increase-aids-cancer-2021/
96% of it all is already PATENTED, for future ‘vaccinations’.. Don’t we have enough DEAD people from the first ‘version’ of the genetically modifying injections???
10/4/2022: The new executive order explained by Todd Callender AND new presumptive PATENTED art of HUMAN SPECIES = Homo Borg Genesis at https://forbiddenknowledgetv.net/todd-callendar-stopping-the-who-camps-medical-tyranny-with-targeted-strategies/ and at https://beforeitsnews.com/alternative/2022/10/homo-borg-genesis-the-new-human-species-explained-by-insurance-ceo-todd-callendar-3781296.html
9/26/2022 : Karen Kingston speaks about more dots pointing to the connections of this new order, NOBODY was asking for or agreed with: https://www.brighteon.com/4d3c05c2-a74c-4de6-958b-6557365c22f5
On Sep 12th 2022 many people were shaken by what they saw in president Biden’s executive order (with no assigned number yet), a quote repeated zillion times, like for example by:
and also in this later one (not being properly linked by sub-stack for ‘some’ reason):
https://joebot.substack.com/p/synthetic-salvation-on-genomics-min
with another complementary article on that topic at: https://www.technologyreview.com/2022/11/16/1063300/billion-dollar-mega-rich-live-forever/
The 100% ANTI-HUMAN, ‘almost’ on the 13th of Sep announced ‘order’ by Biden says, quote:
“We need to develop genetic engineering technologies and techniques to be able to write circuitry for cells and predictably program biology in the same way in which we write software and program computers; unlock the power of biological data, including through computing tools and artificial intelligence; ….”
the entire text is available at:
A very good summary of some of the details of this order can be also found at:
https://www.bitchute.com/video/cE9UkxE5CANO/ as part 1 and its part 2 at:
https://www.bitchute.com/video/T8iMkKymO5Xp/
and just today, 9/25/2022, Dr. Mercola joined the discussion too:
So apparently we are here, on this planet ‘out of chaos’, evolved in a progressive liberal way from apes (few of which strangely are still left out there, possibly for vaccination research purposes), and now, the circuitry of our cells shall be RE-DESIGNED by programmers into PREDICTABLE BIOLOGY?? But as of today, 9/27/2022, even the apes were abandoned, because now the common ancestor of all mammals looks like this:
according to https://phys.org/news/2022-09-reconstruct-genome-common-ancestor-mammals.html, the COMPUTATIONAL RECONSTRUCTION was just published in the prestige PNAS journal, with the first author from UC Davis Genome Center.. Just wonder, who is writing the genomic code for the reconstruction and even more important who is paying for this??? Btw. wouldn’t it be nicer to put the hair little bit more towards the head and trimm some of the overal coverings instead and also add some features implying we deal with male or female???
Back to the WH (white house, which should be green, due to the disappearing oil reserves) plan, so what is wrong with us? Would timing of human bowel movements*(based on Bill Gates involvement with toiletes..) or viral infections belong into that category? Because if so, then the pandemics would be in fact 100% programmable with this ‘plan’.. Here the proof: https://uncutnews.ch/dokumentarfilm-plandemic/
But, wasn’t it the case already? The new executive order also mentions, quote : “The COVID-19 pandemic has demonstrated the vital role of biotechnology and biomanufacturing in developing and producing life-saving diagnostics, therapeutics, and vaccines that protect Americans and the world.“, something clearly very different from what we see:
The andante prelude to the above executive order insanity happened in 2013, with an announcement of the ‘Brain Initiative’ by somebody already with lot of history behind:
https://ugetube.com/watch/the-obama-revolution_wuhponhmhdos4h6.html
The new program was based heavily on ‘Human Genome Project’ and Optogenetics was one of its foundations, followed by Magnetogenetics, both, subjects to remote control via electromagentic fields.
On the 9/19/2019, another Executive Order 13887, featured a plan to control the future influenza pandemics, right on time, without chicken eggs, involving the top national security agencies.. Since when an influenza has to be tight with national security issue, with military involved, with the last pandemic being >100 years ago in the age with the science at totally different level than today??? As it appears the subsequent WARP SPEED for that development was a military contract: https://rumble.com/v1jyxbn-warner-mendenhall.html
Thanks to Mr. Trump, the chickens were saved and the can of reprogramming synthetic biology got opened, leaving no place for the real Creator any more. And the man named Edwin Messe III, who was the Att. General in 1986, in the time of H.R.5546 - National Childhood Vaccine Injury Act of 1986, immediately after the 9/19/2019, got the Medal of Freedom (10/8/2019) from president Trump, all described very well in Dr. Heiko Schoening’s book ‘Game Over’. All these turbulent political facts must be somehow, quietly driven by new facts, from the science community..
So optics is about light, mass-less particles, their generation, propagation and interactions with matter, genetics is about genes, in main science only matter based, and combined together it is about genes/biology driven by light. ‘BRAIN 2025 a scientific vision’, similar to many other futuristic documents with the final dates of specific goals (separate not NIH related example ‘Weather As A Force Multiplier: Owning The Weather In 2025’ http://www.dickatlee.com/issues/environment/climate/mod/pdfs/USAF_weather_force_multiplier.pdf ) was issued by NIH in 2014 with the title: ”Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Working Group Report to the Advisory Committee to the Director, NIH” and specified in their executive summary few points:
-Identify and provide experimental access to the different brain cell types to determine their roles in health and disease. It is within reach to characterize all cell types in the nervous system, and to develop tools to record, mark, and manipulate these precisely defined neurons in the living brain. We envision an integrated, systematic census of neuronal and glial cell types, and new genetic and non‐genetic tools to deliver genes, proteins, and chemicals to cells of interest in non‐human animals and in humans.
-Demonstrating causality: Link brain activity to behavior with precise interventional tools that change neural circuit dynamics. By directly activating and inhibiting populations of neurons, neuroscience is progressing from observation to causation, and much more is possible. To enable the immense potential of circuit manipulation, a new generation of tools for optogenetics, chemogenetics, and biochemical and electromagnetic modulation should be developed for use in animals and eventually in human patients.
-The past decade has seen the development of remarkable genetic tools including calcium indicators (e.g. GCaMP), optogenetic tools (e.g. Channelrhodopsin), synaptic monitors (e.g. SynaptopHluorin), chemogenetic tools (e.g. RASSLs/DREADDs), and a variety of tags that permit proteins to be visualized in vivo. By their nature, using these tools requires the ability to deliver a gene to a neuron or neurons of interest (‘genetic access’).
The gene delivery was envisioned in many ways, quote:
“A full exploration of methods for targeting genes, proteins, and chemicals to specific cell types is highly desirable. One question that should be considered during the initial stages of the BRAIN Initiative is whether genome engineering by conventional transgenesis could be superseded by methods that are faster, cheaper, and more easily generalized across species. Mouse husbandry is slow and expensive, and generating the right multi‐transgenic strains is a financial and temporal drag on the progress of neuroscience research. Furthermore, we wish to study other species as well. The BRAIN Initiative should solicit new ideas for cell‐type specific delivery of transgenes, perhaps based on viruses bearing small specific regulatory regions for intersectional cell‐type definition via (for example) multiple recombinases; or viruses driving efficient, specific integration of exogenous genes into the genome through cutting‐edge tools such as CRISPRs and TALENs; or antibody‐targeted liposomes.“
The manifesto of the Obama’s ‘BRAIN INITIATIVE’ says it all: https://obamawhitehouse.archives.gov/BRAIN
The origin of optogenetics goes back to the first expression of light-driven protein OPSIN (normally only in eyes) in genetically engineered neuronal cells which fire up charges upon illumination. One of the optogenetics founders, Deisseroth, defined this ‘optogenetic emerging opportunity’ as: “the combination of genetic and optical methods to achieve gain or loss of function of well-defined events in specific cells of living tissue”.^1, 2, ,3. The goal was to genetically RE-engineer natural proteins for their maximum output related to LIGHT-CURRENT conversion, an analogy in physics, that’s the corresponding photoelectric effect and its inversion which happens in every light bulb. An extensive resource of the optogenetic tool kits to ‘build better’ the new synthetic biology can be found at: https://web.stanford.edu/group/dlab/optogenetics/ Another source claims optogenetics is ‘just Controlling Neurons with Photons”, https://www.optica-opn.org/home/articles/volume_29/april_2018/features/optogenetics_controlling_neurons_with_photons/
with a small quote from it:“Optogenetic modifications in mice have shown no adverse impacts to their health,” said Rogers, “so it’s conceivable that these approaches could one day be approved for safe use in humans.” Just wonder, did they got the mice to tell the scientists ‘Good morning’ and smile while looking at itself in the mirror..?
This article is also mentioning that:”red light generally inhibits a response in the brain, blue light stimulates one, and green can do either depending on the targeted opsin.” Well there are the light devices which with a specific frequency changes between blue and red lights are capable of healing pain. A good 2 overviews of the so called Low Level Light Therapy (LLLT) can be found at: https://pubmed.ncbi.nlm.nih.gov/24049929/ and at https://pubmed.ncbi.nlm.nih.gov/31405692/
Back to the 2014 Brain Initiative summary, which states: “Emphasis should be placed on achieving modulation of circuits in patterns that mimic natural activity.”
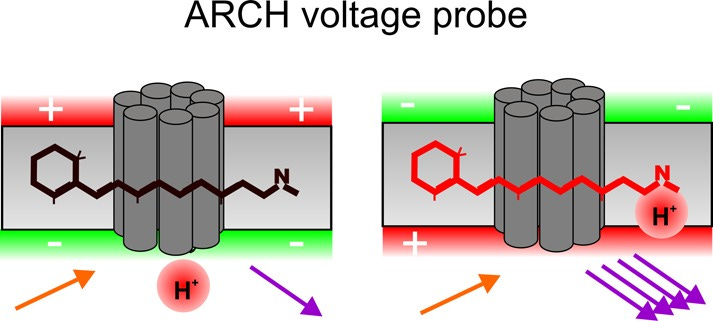
A group of the so called genetically encoded voltage indicators (GEVIs) can be used for mapping neural circuits. A 2012 publication^4 illustrates how the neuronal cell membranes voltage indicators unable imaging of the neuronal circuit operations, which correlate with brain state and behavior. The principal was demonstrated with bacterial opsin proteins binding Vitamin A, which upon light activation lead to charge movement and effectively change the membrane potential:
Engineering different ‘color fluorescing domains’ (frequency) allows for action potentials changing correspondingly with the moving domains.
All the light absorbing features of proteins are dictated by the spacial distribution of aromatic amino acids (tyrosine described as y or Y, tryptophan w or W, phenylalanine f or F), and that is in part dictated by electrostatic interactions of all charged groups within the 3D fold of the protein. The family of membrane anckered proteins consists among others of s.c. GPCR proteins, 7 helix bundle (above picture) which is voltage sensitive, while facing 2 differently charged sides of every membrane, dividing inside-outside of the cell, which we all know is all about ions distribution, movements and polarity (inside +-, outside -+). The current flow accompanying all the motions is always EMF dependent. The trial to DESIGN proteins, which would be giving the strongest voltage signals ended with a patent for so called ARCHONS, seven of them. The owners of that 2019 patent (US 2019/0004032 A1) are MIT and NIH and the title:”Genetically encoded red fluorescent voltage sensors enabling millivolt-resolution and high-speed neural voltage imaging.” The distribution of the aromatic amino acids in one of the archons, in SARS-CoV2 Spike (expressed in all the injected covid victims) and in part of one of the largest human proteins, titin, in a simple linear representation of their entire aromatic amino acid sequence (aa’s) is illustrated below, with unfortunately changed format, in which the point I wanted to make, is no more clearly ‘visible’, but at least a try:
archon1 homolog at https://www.ncbi.nlm.nih.gov/protein/QHY11315.1
1 y w fyf w yy
61 y ff yy y w f
121 y ww f y
181 f w y w f f f
241
in the entire SARS-CoV2 Spike:
1 f f y f yy f f ff
61 wf f f yf w f
121 f f f yy w f y f y f
181 f f f y f y f f
241 y w yy y f y
301 f y f f f f f y w
361 y y f f y f y f
421 y y f w y y y f f y
481 f yf y f y y f
541 f f f f f f
601 y w y f y
661 y y y f
721 y y f
781 f y f f f f f f y
841 f y w f f
901 y f y f
961 f y
1021 f y f f y f
1081 f f wf fy f y
1141 f yf
1201 y y w wy w f f
1261 y
and in the first ~1200 residues of the largest human protein, titin, just for a comparison:
1 f f f wf f
61 y f
121 fy f y y y
181 f
241
301 w y
361 w y y
421
481 f
541
601
661 y y y
721 f
781 y f f
841 y f
901 y w y y f f f
961 f y f f f
1021 yf f yw y
1081 y y f y y
1141 y f f y y y y f
1201 f y f y
1261 f y wy y f y f
The difference in densities of the UV light absorbing aromatic essential aa’s tryptophan, phenylalanine and tyrosine, can also be expressed in numbers:
archon : y(11) , f(10), w(7) in 248 residues <=synthetics
Spike : y(54), f(77), w(12) in 1273residues <=synthetics
titin : y(34), f(30), w(6) in 1280 in residues <=natural
Here examples of some essential human proteins with their % content of aromatic aa’s in the entire protein, starting with the PATENTED SARS-CoV-2 Spike and the archons:
Spike2020 with 1273 aa’s, total aromatic aa’s 11.1%
Phe (F) 77 6.0%
Trp (W) 12 0.9%
Tyr (Y) 54 4.2%
all the seven patented archon proteins have the same content of aromatic aa’s total 11.1% AND their spacial 3D distribution is CONSERVED:
Phe (F) 10 4.0%
Trp (W) 7 2.8%
Tyr (Y) 11 4.3%
Long-wave-sensitive opsin 1 (Human Red cone photoreceptor pigment), uniprot code P04000.2, total 13.7%:
Phe (F) 20 5.5%
Trp (W) 14 3.8% Tyr (Y) 16 4.4%
Short-wave-sensitive opsin 1 (Blue cone photoreceptor pigment) (Blue-sensitive opsin) (BOP, uniprot code P03999), total 16.1%
Phe (F) 33 9.5%
Trp (W) 7 2.0% Tyr (Y) 16 4.6%
HEMOGLOBIN , subunit delta (Delta-globin) HBD_HUMAN (P02042) 147 aa’s, total 9%
Phe (F) 8 5.5%
Trp (W) 2 1.4%
Tyr (Y) 3 2.1%
ALBUMIN_precursor. HUMAN (P02768), 606 aa’s, total 9.2%
Phe (F) 35 5.8%
Trp (W) 2 0.3%
Tyr (Y) 19 3.1%
MYO1E_HUMAN (Q12965), human myosin 1108 aa’s, total 10%
Phe (F) 49 4.4%
Trp (W) 12 1.1%
Tyr (Y) 50 4.5%
TITIN_HUMAN (Q8WZ42) Titin (Connectin), largest human protein with 34350 aa’s, total 6.9%
Phe (F) 908 2.6%
Trp (W) 466 1.4%
Tyr (Y) 999 2.9%
KERATIN HUMAN, 472 aa’s, total 6%
Phe (F) 12 2.5%
Trp (W) 2 0.4%
Tyr (Y) 14 3.0%
ELASTIN human, 786 aa’s, total 3.9%
Phe (F) 16 2.0%
Trp (W) 0 0.0% !!!
Tyr (Y) 15 1.9%
Collagen alpha-1(VII) chain · Q02388, 2944 aa’s, total 2.5%
Phe (F) 39 1.3%
Trp (W) 19 0.6%
Tyr (Y) 36 1.2%
Actin human (P60709), cytoplasmic 1, 375 aa’s, total 8.6%
Phe (F) 13 3.5%
Trp (W) 4 1.1%
Tyr (Y) 15 4.0%
Fibronectin (Human), P02751, 2477 amino acids, total 8%
Phe (F) 54 2.2%
Trp (W) 40 1.6%
Tyr (Y) 104 4.2%
Coagulation factor VIII (Human) 2351 aa’s, total 9.6%
Phe (F) 109 4.6%
Trp (W) 37 1.6%
Tyr (Y) 79 3.4%
Coagulation factor XIII A chain, (Human), 732 aa’s, total 10.4%
Phe (F) 32 4.4%
Trp (W) 15 2.0%
Tyr (Y) 29 4.0%
Cadherin-3 (Human), 829 aa’s, total 6.9%
Phe (F) 26 3.1%
Trp (W) 8 1.0%
Tyr (Y) 23 2.8%
Integrin alpha-5 · Homo sapiens (Human), P08648, 1049 aa’s, total 9.5%
Phe (F) 47 4.5%
Trp (W) 13 1.2%
Tyr (Y) 38 3.6%
Cystic fibrosis transmembrane conductance regulator (Human), P13569, 1480 aa’s, total 10%
Phe (F) 85 5.7%
Trp (W) 23 1.6%
Tyr (Y) 40 2.7%
Acetylcholine receptor subunit gamma (Human), P07510, 517 aa’s, total 8.6%
Phe (F) 21 4.2%
Trp (W) 12 2.4%
Tyr (Y) 10 2.0%
ACE2 receptor (Human), Q9BYF1, 805 aa’s, total 11.8%
Phe (F) 39 4.8%
Trp (W) 23 2.9%
Tyr (Y) 33 4.1%
These numbers for all the different types of proteins from different tissues, seem to indicate one, the highest amount of aromatic amino acids is among the proteins which deal with VISION (photons within very narrow wavelength range ~350-750nm, equivalent with ~400THz to ~700THz frequency ranges), with the exception of ACE2 receptor... And the PATENTED Spike2020, is among them.. The very special motif WxWYxW in Spike having its homolog within the in 1991 by Biogen, Inc. (Cambridge, MA) filed patented HIV surface glycoprotein gp160 protein, US 5,871,732 “Anti-CD4 antibody homologs useful in prophylaxis and treatment of AIDS, ARC and HIV infection“ (link to it from https://www.ncbi.nlm.nih.gov/protein/AAE05938.1 ), is aligned automatically by BLAST software the following way:
20.4 bits(41) 0.75 Compositional matrix adjust. 8/27(30%) 13/27(48%) 1/27(3%)
Query 6 KYQHLWRWGWR-WGTMLLGMLMICSAT 31 HIV-1 gp160
KY+ +W+W W + G++ I T
Sbjct 1205 KYEQYIKWPWYIWLGFIAGLIAIVMVT 1231 SARS-CoV-2 Spike
also the very same motif has its homolog in the ACE2 receptor itself!
Mutation of the 169Arg causes 95% loss of angiotensin I cleavage, also that residue binds chloride
*
DYNE-RL-WAWESWRSEVGKQL ACE2
Y E + W+W W + L
KY-EQYIKWPWYIWLGFIA-GL SARS-CoV-2The extremely disturbing fact, clear from the 2018 R.Baric’s 9884895-B2 patent (https://www.ncbi.nlm.nih.gov/protein/AVY23063.1) on “Methods and compositions for chimeric coronavirus spike proteins“ is that this motif is actually present in the Spike of :
-2005 Transmissible gastroenteritis virus (NIH entry CAB91145.1) with the sequence alignment of the SARS-CoV-2 SPike 2020:
350 bits(897) 3e-104 Compositional matrix adjust. 259/803(32%) 393/803(48%)
Que1339 NLTGEIDDLEFRSEKLHNTTVELAILIDNINNTLVNLEWLNRIETYVKWPWYVWL
N+ EID L E+A N+N +L++L+ L + E Y+KWPWY+WL
Sbj1178 NIQKEIDRLN-----------EVA---KNLNESLIDLQELGKYEQYIKWPWYIWL
Query 1396 --LIGLVVIFCIPLLLFCCCSTGCCGCI 1419
+ GL+ I + ++L CC T CC C+
Sbjct 1223 GFIAGLIAIVMVTIML--CCMTSCCSCL 1244
-2016 mouse hepatitis virus, whose spike (NIH entry NP_045300.1 )is almost 40% identical within ~70% sequence coverage with Spike 2020 (Sb line below)
483 bits(1242) 2e-152 Compositional matrix adjust. 277/749(37%) 413/749(55%)
Qu1208 PDLSL-DFEKLNVTLLDLTYEMNRIQDAIKKLNESYINLKEVGTYEMYVKWPWYVW
PD+ L D +N +++++ E++R+ + K LNES I+L+E+G YE Y+KWPWY+W
Sb1157 PDVDLGDISGINASVVNIQKEIDRLNEVAKNLNESLIDLQELGKYEQYIKWPWYIW 1214
Query 1264 LLIGLAGVAVCVLLFFICCCTGCGSC 1295
L +A+ ++ +CC T C SC
Sbjct 1212 LGFIAGLIAIVMVTIMLCCMTSCCSC 1243-2005 Avian infectious bronchitis virus isolate IBV-p65 (NIH entry AAY24433.1)
359 bits(921) 6e-109 Compositional matrix adjust. 200/548(36%) 296/548(54%) 22/548(4%)
Qu1056 ILDIDSEIDRIQGVIQGLNDSLIDLEKLSILKTYIKWPWYVWLAIAFATIIFILIL
+++I EIDR++ V + LN+SLIDL++L + YIKWPWY+WL+ I ++++
Sb1176 VVNIQKEIDRLNEVAKNLNESLIDLQELGKYEQYIKWPWYIWLGFIAGLIAIVMVT
Query 1112 GWVFFMTGCCGC 1123
++ MT CC C
Sbjct 1232 IMLCCMTSCCSC 1243-2006 porcine epidemic diarrhea virus, (NIH entry CAA80971.1 )
332 bits(851) 3e-98 Compositional matrix adjust. 237/775(31%) 363/775(46%)
Qu1300 LINNINNTLVDLEWLNRVETYIKWPWWVWLIIVIVLIFVVSLLVFCCISTGCCGC
+ N+N +L+DL+ L + E YIKWPW++WL + LI +V + + C T CC C
Sb1189 VAKNLNESLIDLQELGKYEQYIKWPWYIWLGFIAGLIAIVMVTIMLCCMTSCCSC -2009 Porcine hemagglutinating encephalomyelitis virus (NIH entry QTF73995.1 )
Qu1231 SLDYINVTFLDLQDEMNRLQEAIKVLNQSYINLKDIGTYEYYVKWPWYVWL 1298
+ IN + +++Q E++RL E K LN+S I+L+++G YE Y+KWPWY+WL
Sb1150 DISGINASVVNIQKEIDRLNEVAKNLNESLIDLQELGKYEQYIKWPWYIWL 1218
Query 1299 LIGLAGVAMLVLLFFICCCTGC 1320
+A++++ +CC T C
Sbjct 1219 GFIAGLIAIVMVTIMLCCMTSC 1240- And in 1987 (!!!) Feline infectious peritonitis virus (FIPV), its Spike, called E2 peplomer glycoprotein (NIH entry CAA29535.1), ~32% IDENTICAL with Spike2020
344 bits(882) 4e-102 Compositional matrix adjust. 253/801(32%) 388/801(48%)
Qu1346 EIDDLEFRSEKLHNTTVELAILIDNINNTLVNLEWLNRIETYVKWPWYVWL 1403
EID L E+A N+N +L++L+ L + E Y+KWPWY+WL
Sb1180 EIDRLN-----------EVA---KNLNESLIDLQELGKYEQYIKWPWYIWL 1225
Query 1397 --LIGLVVVFCIPLLLFCCFSTGCCGCI 1424
+ GL+ + + ++L CC T CC C+
Sbjct 1219 GFIAGLIAIVMVTIML-CCM-TSCCSCL 1244This 1998 entry, despite of having ~1400 aa’s, slightly more than Spike 2020, the entire N-terminal domain of the first ~600 amino acids, is ‘just not aligning’ by the NIH’s BLAST, thus the 32% identity is only for that substantially cut Spike with its S2 domain only. Even cutting the S2 domain from Spike and trying to automatically align it with the entire 1987 peplomer glycoprotein, just does not work… Thus by hand, the first line with NO shifts, the first 70 amino acids of both N-terminals (i.e. S1 domains):
MIVLVTCLLLLCSYHTVLSTTNNECIQVNVTQLAGNENLIRDFLFSNFKEEGSVVVGGYYPTE
M V + L L+ S L+T + + G + F S + + + +
MFVFLVLLPLVSSQCVNLTTRTQLPPAYTNSFTRGVYYPDKVFRSSVLHSTQDLFLPFFSNVT
VWYNCSR <<<1987 (FIPV) peplomer glycoprotein
W+ +
-WFHAIHV <<<Spike 2020
The entry https://www.ncbi.nlm.nih.gov/protein/CAA29535.1, pointing to the 1987 publication^16, contains description of the FURIN cleavage site, quote:
Site order(958,962)
/note="S2' furin cleavage site"And here the description of the 2020 Spike’ entry https://www.ncbi.nlm.nih.gov/protein/1796318598 furin site :
Site 685..686
/note="S1/S2 furin cleavage site"One of the by Baric’s used chimeric coronavirus spike proteins in his 2018 patent lists the strain SHC014-CoV. Search for similarities of its spike proteins to Spike2020 leads to this construct “receptor binding domain/SpyTag003 fusion protein“, almost identical to Spike 2020, overlapping exactly its ACE2 receptor binding portion…
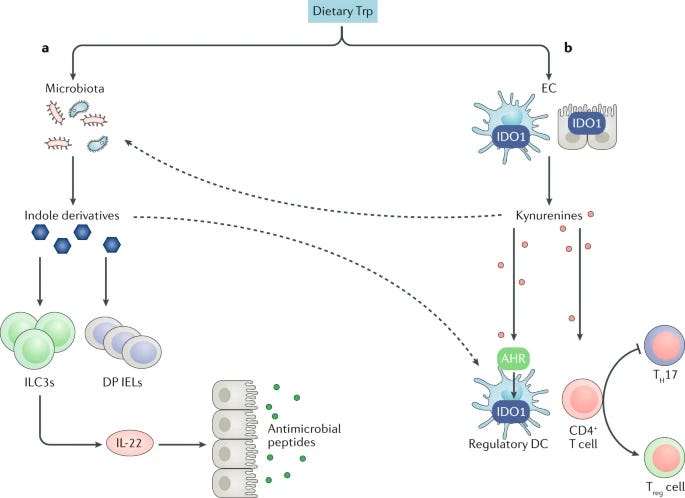
Strangely there is also growing amount of new literature about graphene and aromatic amino acids, like tryptophan or tyrosine, for example ^6,^7, ^8, ^9, ^10, taking just the tryptophan alone. And on the other hand there are publications which indicate tryptophan involvement in cancer ^11 and according to Nature article, SO much more ^12, quote: “Together, these functions suggest that, during evolution(??<-my addition), Trp (i.e. tryptophan) metabolism has become part of the cellular and organismal communication strategies that align food availability with physiology and behaviour.” According to that article it is also essential in immune and auto-immune response:
The very same can be found when looking for tyrosin importance in the entire biology.
The chirality of all the amino acids in nature is essential, the proteins are always containing the L-form of every amino acid in their sequences, never their MIRRORED IMAGE in the D-form which would rotate the circular light in the opposite direction. Aren’t microwaves capable of switching that chirality L→D, thus making the amino acid useless, or even toxic? Yes, the one of ~2.4GHz..?
The chemical bond called pi-pi stacking of the large flat electron rings in tryptophan with other molecules with the same characteristics, like genetic material DNA or RNA, makes it’s special features even more important, like direct interaction with DNA^13, small peptides like KWGK (present in one of the SARS-CoV2 orf proteins) once bound to G or C polynucleotides, can switch completely the chirality of surrounding DNA^14. The abstract of that publication also states “This suggests that tryptophan intercalation may act as a discriminating factor in recognizing Z and B-forms and may have a potential role in Protein-Nucleic acid interactions that are important for transcription.“ Another very important feature involving tryptophan is that their aromatic π–π stacking interactions play a key role in directional control of self-assembly^15, but so does the RGD (Arginine-Glycine-Aspartic Acid, tripeptide) motif, present only in SARS-CoV-2 Spike and no other coronaviruses:
Coming back to the SARS-CoV-2 Spike opsin issues, a BLAST (NIH bioinformatics software) comparison with the Long-wave-sensitive opsin 1 (LWS) gives astonishing 50% of idenitities for ~75% sequence coverage when run with very customized parameters, looking more in detail at shorter fragments (which are extremely important for building epitopes), here some examples of the output:
24.0 bits(49) 0.026 9/18(50%) 9/18(50%) 9/18(50%)
Query 221 IVLMVTCCIIPLAIIMLC 238 Opsin 1 (LWS)
IV MVT IMLC << IDENTITIES
Sbjct 1227 IV-MVT--------IMLC 1235 Spike 2020
Query 30 YTNSNSTRGPFEGPNYHIAPRWVY 53 Opsin 1 (LWS)
YTNS TRG VY
Sbjct 28 YTNS-FTRG-------------VY 37 Spike 2020
Query 306 LP----AYFA---KSATI 316 Opsin 1 (LWS)
LP YFA KS I << IDENTITIES
Sbjct 84 LPFNDGVYFASTEKSNII 101 Spike 2020
Query 307 PAY---FAKSATIYNPVIY---VFMNRQFRNCILQ----LF 337
PAY F++ +Y Y VF R +L+ LF
Sbjct 26 PAYTNSFTRG--VY----YPDKVF-----RSSVLHSTQDLF 55
Query 249 VAKQQKES 256
VAK ES
Sbjct 1189 VAKNLNES 1196
Query 225 VTCCIIPLAI 234
V C +P+AI
Sbjct 615 VNCTEVPVAI 624
Query 97 VADLAETVIASTIS 110
VA + ++IA T S
Sbjct 687 VA--SQSIIAYTMS 698
Query 135 LCGITGLWSLAIISWERWMVVCKPFGNVRFDA 166
LC PFG V F+A
Sbjct 335 LC---------------------PFGEV-FNA 344
Query 305 ALPAYFAKSAT 315
A+P+ F+ S T
Sbjct 713 AIPTNFTISVT 723
Query 106 ASTIS----IVNQ 114
AS++ +VNQ
Sbjct 942 ASALGKLQDVVNQ 954
Query 337 FGKKVDDGSELSSASKTE 354
FG +D SKT+
Sbjct 106 FGTTLD--------SKTQ 115
Query 28 FT--YTNSNSTRG 38
FT Y++S +RG
Sbjct 392 FTNVYADSFVIRG 404
Query 328 QFRNCILQLFGKKVDDGSELSS-AS 351
QF I G K+ D LSS AS
Sbjct 926 QFNSAI----G-KIQDS--LSSTAS 943
Query 201 TSC-GPDVFSGSSY-PGVQSY 219
T C G + F Y P +QSY
Sbjct 478 TPCNGVEGF--NCYFP-LQSY 495
and since certain motifs are 'not found' by the NIH BLAST, a comparison done by hand
FS-WIWAAVWTAPPIFGW Opsin 1 (LWS)
+ W W +W + I G
YIKWPWY-IWLG-FIAGL Spike 2020
and again, many, many more fragments. The comparison to the short wavelength sensitive (SWS) opsin results in different homologous sections, for example:
24.1 bits(48) 0.023 10/20(50%) 12/20(60%) 4/20(20%)
Query 221 TQLLRALKAVAAQQ----QE 236 Opsin (SWS)
TQL RAL ++A +Q QE
Sbjct 761 TQLNRALTGIAVEQDKNTQE 780 Spike2020
Query 8 EFYLFKNI 15 Opsin (SWS)
EF FKNI
Sbjct 191 EF-VFKNI 197 Spike 2020
Query 182 CSCG 185 Opsin (SWS)
CSCG
Sbjct 1248 CSCG 1251 Spike2020
Query 89 FP---VFVASCNG--YFV 101
FP VFV+ NG +FV
Sbjct 1089 FPREGVFVS--NGTHWFV 1104
Query 207 FCFIVPLSLI 216
F F+V L L+
Sbjct 2 FVFLVLLPLV 11
Query 67 QPLNYILVNVSF 78
QP Y +V +SF
Sbjct 506 QP--YRVVVLSF 515
Query 113 LGTVAGLVTGWSLAFLAFERYIV 135 Opsin (SWS)
LG +AGL++ IV
Sbjct 1218 LGFIAGLIA------------IV 1228 Spike2020
Query 125 LAFLAFER 132
L F F R
Sbjct 560 LPFQQFGR 567In the short-wavelength sensitive (SWS) opsin there is no tryptopan cluster ’WIWAAVWTAPP’ like in the LWS version, thus the presence of homology of this cluster in Spike 2020 (aligned by hand in the previous tab), speaks more for a probability of Spike being ‘related’ to the long wavelength (red) opsin, involved among others, in colorblindness. How similar is this human LWS opsin with the by Dr. Walensky patented 2 sequences at https://patents.google.com/patent/US9290545B2/en completely embedded in SARS-CoV-2 SPike??? NIH BLAST gives NO OVERLAP, no matter what you do, and here by hand alignment with almost the entire length with one of the Walensky’s heptad repeats, with the first line being the patent, second the identities with the sequence of the LWS protein in the third line:
NVLYENQKQIANQFNKAI---SQIQESLTTTSTALGK-LQDVVNQNAQALNTLVKQLSSNFGA-
Y K +N I + Q + GK + D + + +LSS +
APAYFA-KSATI-YNPVIYVFMNRQFRNCILQLF-GKKVDD-------G--S---ELSSASKT
-ISSVLNDILSRLDKVE-AE <<< Walensky's patent
+SSV S V A <<< IDENTITIES
EVSSV-----SS---VSPA <<< HUMAN LWS opsin 1Does it look RANDOM?? BLAST does think so, I do not.
The case of magnetogenetics was covered to some extend in the previous post of mine:
but there is so much more to it, that it needs more consideration in the next post.
The at the begin of this post mentioned changed bloody system, related to a 2521 amino acid long protein named PIEZO1, can possibly help to explain the huge clots in human bodies (deceased and alive) alone due to the properties of this sensor. Clearly genetic reprograming of that sensor must affect its functions, which according to uniprot server (complete biochemical characterization of known proteins) are, quote:
“Pore-forming subunit of a mechanosensitive non-specific cation channel (PubMed:23479567, PubMed:23695678). Generates currents characterized by a linear current-voltage relationship that are sensitive to ruthenium red and gadolinium. Plays a key role in epithelial cell adhesion by maintaining integrin activation through R-Ras recruitment to the ER, most probably in its activated state, and subsequent stimulation of calpain signaling (PubMed:20016066). In the kidney, may contribute to the detection of intraluminal pressure changes and to urine flow sensing. Acts as shear-stress sensor that promotes endothelial cell organization and alignment in the direction of blood flow through calpain activation (PubMed:25119035).
Plays a key role in blood vessel formation and vascular structure in both development and adult physiology (By similarity).
Acts as sensor of phosphatidylserine (PS) flipping at the plasma membrane and governs morphogenesis of muscle cells. In myoblasts, flippase-mediated PS enrichment at the inner leaflet of plasma membrane triggers channel activation and Ca2+ influx followed by Rho GTPases signal transduction, leading to assembly of cortical actomyosin fibers and myotube formation“
The universal presence of graphene compounds in our air, in water, in our bodies, possibly in many covid injection materials, begs the question: what to do about it? First would be to strongly regulate the entire production industry of this lethal compound, but also to implement materials, as simple as HYDROGEN PEROXIDE for graphene sheets decomposition^16. The NIST publication describes degradation of the graphene family nano’s (GFN) by simple HYDROGEN PEROXIDE. Here 2 quotes from both publications (repeat from my post
:
“Degradation of graphene by H2O2 at physiologically and environmentally relevant concentrations (1–10 000 × 10−6m) is reported. Exposure to H2O2 leads to the formation of holes on graphene sheets. The degradation phenomenon is dependent on concentration of H2O2 and incubation time.“
“GFNs in the natural environment are also subject to chemical transformation by strong, naturally occurring oxidizing agents such as hydrogen peroxide (H2O2), found in rain and natural waters. The degradation of graphene (and most likely, other GFNs) can occur at concentrations of H2O2 that naturally occur in surface waters (51 mg/L to 231 mg/L or 1 x 10−3 mol/L to 7 × 10−3 mol/L), leaving defects on the surface of the GFN. Similarly, iron/H2O2 driven Fenton chemistry (with or without UV irradiation), a treatment technique commonly applied in wastewater treatment plants, can generate reactive oxygen species (ROS; such as hydroxyl radical, •OH) which can cause defects in the GFN structure, and even lead to complete degradation to CO2 at sufficiently high concentrations of the reactants.“
Last but not least, the post:
mentions binding of the toxic SARS-CoV-2 Spike by the IVERMECTIN. But it is known that IVERMECTIN equally binds GRAPHENE^17, or actually more frequently one will find publications describing detection of IVERMECTIN using grapehne compounds, which implies, they both need to bind…. For those, who do not believe in in the viral existence that video will be a good argument: https://www.instagram.com/tv/CjWbootgI9A/
Literature.
Deisseroth, K. (2010). Controlling the brain with light. Scientific American, 303(5), 48–55.
Deisseroth, K. (2011). Optogenetics. Nature Methods, 8(1), 26–29.
Deisseroth, K., Feng, G., et al. (2006). Next-generation optical technologies for illuminating genetically targeted brain circuits. The Journal of Neuroscience, 26(41), 10380–10386.
Hiroki Mutoh et al. “Genetically Engineered Fluorescent Voltage Reporters” ACS Chem. Neurosci. 2012, 3, 585−592.
Bumjun et al. “A dimeric fluorescent protein yields a bright, red-shifted GEVI capable of population signals in brain slice. Scientific Reports (2018) 8:15199.
“Tyrosine and Tryptophan Have Different Binding Sites on Graphene Oxide”
January 2016 New Physics Sae Mulli 66(1):103-107.
https://www.researchgate.net/publication/292176040_Tyrosine_and_Tryptophan_Have_Different_Binding_Sites_on_Graphene_Oxide
“Investigation of L-Tryptophan Electrochemical Oxidation with a Graphene-Modified Electrode“ Biosensors (Basel). 2021 Jan 28;11(2):36.
doi: 10.3390/bios11020036.
“Tryptophan-functionalized graphene quantum dots with enhanced curcumin loading capacity and pH-sensitive release“. Journal of Drug Delivery Science and Technology Volume 61, February 2021, 102137 https://www.sciencedirect.com/science/article/abs/pii/S177322472031426X
“Water dispersible graphene noncovalently functionalized with tryptophan and its poly(vinyl alcohol) nanocomposite” Composites Part B: Engineering
Volume 42, Issue 8, December 2011, Pages 2130-2135, at https://www.sciencedirect.com/science/article/abs/pii/S1359836811002265
“Enantioanalysis of tryptophan in whole blood samples using stochastic sensors-A screening test for gastric cancer“ Chirality. 2020 Feb;32(2):215-222.
doi: 10.1002/chir.23155. Epub 2019 Nov 20.
“Tryptophan catabolism in cancer: beyond IDO and tryptophan depletion“ https://pubmed.ncbi.nlm.nih.gov/23090118/
“Tryptophan metabolism as a common therapeutic target in cancer, neurodegeneration and beyond“ https://www.nature.com/articles/s41573-019-0016-5
“Interaction between tryptophan-Sm(III) complex and DNA with the use of a acridine orange dye fluorophor probe. “ https://pubmed.ncbi.nlm.nih.gov/26016416/
“Tryptophan intercalation in G, C containing polynucleotides: Z to B conversion of poly [d(G-5M C)] in low salt induced by a tetra peptide“ J Biomol Struct Dyn. 1996 Aug;14(1):25-30. doi: 10.1080/07391102.1996.10508926.
“Peptide-Based Materials via Molecular Self-Assembly” Chap. 3 by Rein V. Ulijn in ‘Peptide Design Strategies’ 2012.
David G. Goodwin et al. “Detection and Quantification of Graphene Family Nanomaterials in the Environment“ 2018 NIST publication.
M. Mahnashi et al. “Ivermectin detection using Ag@ B, S co-doped reduced graphene oxide nanohybrid”; ToobaRezazadeh et al. “Investigation of adsorption performance of graphene oxide/polyaniline reinforced hollow fiber membrane for preconcentration of Ivermectin in some environmental samples“;
* https://www.thegatewaypundit.com/2022/09/go-bill-gates-samsung-develop-prototype-reinvented-toilet-turns-poop-ashes-fully-recycles-water/